 View on TensorFlow.org
View on TensorFlow.org
|
 Run in Google Colab
Run in Google Colab
|
 View source on GitHub
View source on GitHub
|
 Download notebook Download notebook
|
This tutorial introduces autoencoders with three examples: the basics, image denoising, and anomaly detection.
An autoencoder is a special type of neural network that is trained to copy its input to its output. For example, given an image of a handwritten digit, an autoencoder first encodes the image into a lower dimensional latent representation, then decodes the latent representation back to an image. An autoencoder learns to compress the data while minimizing the reconstruction error.
To learn more about autoencoders, please consider reading chapter 14 from Deep Learning by Ian Goodfellow, Yoshua Bengio, and Aaron Courville.
Import TensorFlow and other libraries
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import tensorflow as tf
from sklearn.metrics import accuracy_score, precision_score, recall_score
from sklearn.model_selection import train_test_split
from tensorflow.keras import layers, losses
from tensorflow.keras.datasets import fashion_mnist
from tensorflow.keras.models import Model
Load the dataset
To start, you will train the basic autoencoder using the Fashion MNIST dataset. Each image in this dataset is 28x28 pixels.
(x_train, _), (x_test, _) = fashion_mnist.load_data()
x_train = x_train.astype('float32') / 255.
x_test = x_test.astype('float32') / 255.
print (x_train.shape)
print (x_test.shape)
Downloading data from https://storage.googleapis.com/tensorflow/tf-keras-datasets/train-labels-idx1-ubyte.gz 29515/29515 ━━━━━━━━━━━━━━━━━━━━ 0s 0us/step Downloading data from https://storage.googleapis.com/tensorflow/tf-keras-datasets/train-images-idx3-ubyte.gz 26421880/26421880 ━━━━━━━━━━━━━━━━━━━━ 0s 0us/step Downloading data from https://storage.googleapis.com/tensorflow/tf-keras-datasets/t10k-labels-idx1-ubyte.gz 5148/5148 ━━━━━━━━━━━━━━━━━━━━ 0s 0us/step Downloading data from https://storage.googleapis.com/tensorflow/tf-keras-datasets/t10k-images-idx3-ubyte.gz 4422102/4422102 ━━━━━━━━━━━━━━━━━━━━ 0s 0us/step (60000, 28, 28) (10000, 28, 28)
First example: Basic autoencoder

Define an autoencoder with two Dense layers: an encoder, which compresses the images into a 64 dimensional latent vector, and a decoder, that reconstructs the original image from the latent space.
To define your model, use the Keras Model Subclassing API.
class Autoencoder(Model):
def __init__(self, latent_dim, shape):
super(Autoencoder, self).__init__()
self.latent_dim = latent_dim
self.shape = shape
self.encoder = tf.keras.Sequential([
layers.Flatten(),
layers.Dense(latent_dim, activation='relu'),
])
self.decoder = tf.keras.Sequential([
layers.Dense(tf.math.reduce_prod(shape).numpy(), activation='sigmoid'),
layers.Reshape(shape)
])
def call(self, x):
encoded = self.encoder(x)
decoded = self.decoder(encoded)
return decoded
shape = x_test.shape[1:]
latent_dim = 64
autoencoder = Autoencoder(latent_dim, shape)
autoencoder.compile(optimizer='adam', loss=losses.MeanSquaredError())
Train the model using x_train as both the input and the target. The encoder will learn to compress the dataset from 784 dimensions to the latent space, and the decoder will learn to reconstruct the original images.
.
autoencoder.fit(x_train, x_train,
epochs=10,
shuffle=True,
validation_data=(x_test, x_test))
Epoch 1/10 WARNING: All log messages before absl::InitializeLog() is called are written to STDERR I0000 00:00:1713230807.172583 14227 service.cc:145] XLA service 0x7fe4cc007e20 initialized for platform CUDA (this does not guarantee that XLA will be used). Devices: I0000 00:00:1713230807.172638 14227 service.cc:153] StreamExecutor device (0): Tesla T4, Compute Capability 7.5 I0000 00:00:1713230807.172643 14227 service.cc:153] StreamExecutor device (1): Tesla T4, Compute Capability 7.5 I0000 00:00:1713230807.172646 14227 service.cc:153] StreamExecutor device (2): Tesla T4, Compute Capability 7.5 I0000 00:00:1713230807.172648 14227 service.cc:153] StreamExecutor device (3): Tesla T4, Compute Capability 7.5 131/1875 ━━━━━━━━━━━━━━━━━━━━ 2s 1ms/step - loss: 0.1034 I0000 00:00:1713230807.858362 14227 device_compiler.h:188] Compiled cluster using XLA! This line is logged at most once for the lifetime of the process. 1875/1875 ━━━━━━━━━━━━━━━━━━━━ 5s 2ms/step - loss: 0.0397 - val_loss: 0.0134 Epoch 2/10 1875/1875 ━━━━━━━━━━━━━━━━━━━━ 2s 1ms/step - loss: 0.0124 - val_loss: 0.0111 Epoch 3/10 1875/1875 ━━━━━━━━━━━━━━━━━━━━ 3s 1ms/step - loss: 0.0102 - val_loss: 0.0097 Epoch 4/10 1875/1875 ━━━━━━━━━━━━━━━━━━━━ 2s 1ms/step - loss: 0.0095 - val_loss: 0.0094 Epoch 5/10 1875/1875 ━━━━━━━━━━━━━━━━━━━━ 2s 1ms/step - loss: 0.0092 - val_loss: 0.0092 Epoch 6/10 1875/1875 ━━━━━━━━━━━━━━━━━━━━ 3s 1ms/step - loss: 0.0090 - val_loss: 0.0090 Epoch 7/10 1875/1875 ━━━━━━━━━━━━━━━━━━━━ 2s 1ms/step - loss: 0.0089 - val_loss: 0.0089 Epoch 8/10 1875/1875 ━━━━━━━━━━━━━━━━━━━━ 2s 1ms/step - loss: 0.0088 - val_loss: 0.0090 Epoch 9/10 1875/1875 ━━━━━━━━━━━━━━━━━━━━ 2s 1ms/step - loss: 0.0087 - val_loss: 0.0089 Epoch 10/10 1875/1875 ━━━━━━━━━━━━━━━━━━━━ 2s 1ms/step - loss: 0.0087 - val_loss: 0.0088 <keras.src.callbacks.history.History at 0x7fe6854d9b50>
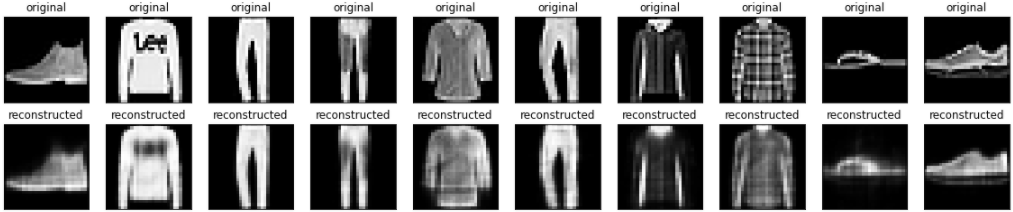
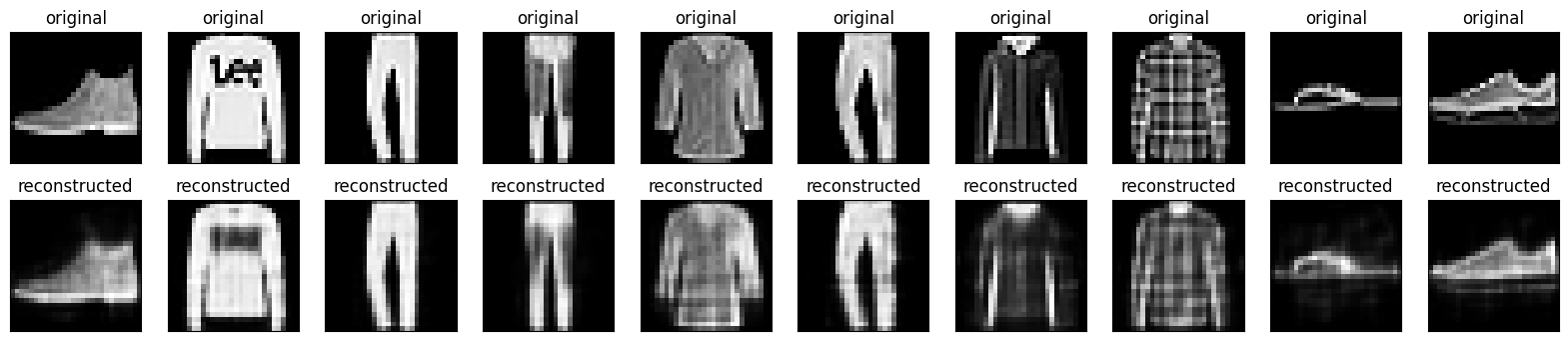
Now that the model is trained, let's test it by encoding and decoding images from the test set.
encoded_imgs = autoencoder.encoder(x_test).numpy()
decoded_imgs = autoencoder.decoder(encoded_imgs).numpy()
n = 10
plt.figure(figsize=(20, 4))
for i in range(n):
# display original
ax = plt.subplot(2, n, i + 1)
plt.imshow(x_test[i])
plt.title("original")
plt.gray()
ax.get_xaxis().set_visible(False)
ax.get_yaxis().set_visible(False)
# display reconstruction
ax = plt.subplot(2, n, i + 1 + n)
plt.imshow(decoded_imgs[i])
plt.title("reconstructed")
plt.gray()
ax.get_xaxis().set_visible(False)
ax.get_yaxis().set_visible(False)
plt.show()

Second example: Image denoising
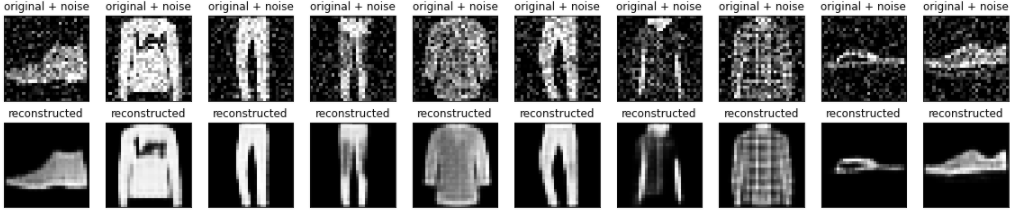

An autoencoder can also be trained to remove noise from images. In the following section, you will create a noisy version of the Fashion MNIST dataset by applying random noise to each image. You will then train an autoencoder using the noisy image as input, and the original image as the target.
Let's reimport the dataset to omit the modifications made earlier.
(x_train, _), (x_test, _) = fashion_mnist.load_data()
x_train = x_train.astype('float32') / 255.
x_test = x_test.astype('float32') / 255.
x_train = x_train[..., tf.newaxis]
x_test = x_test[..., tf.newaxis]
print(x_train.shape)
(60000, 28, 28, 1)
Adding random noise to the images
noise_factor = 0.2
x_train_noisy = x_train + noise_factor * tf.random.normal(shape=x_train.shape)
x_test_noisy = x_test + noise_factor * tf.random.normal(shape=x_test.shape)
x_train_noisy = tf.clip_by_value(x_train_noisy, clip_value_min=0., clip_value_max=1.)
x_test_noisy = tf.clip_by_value(x_test_noisy, clip_value_min=0., clip_value_max=1.)
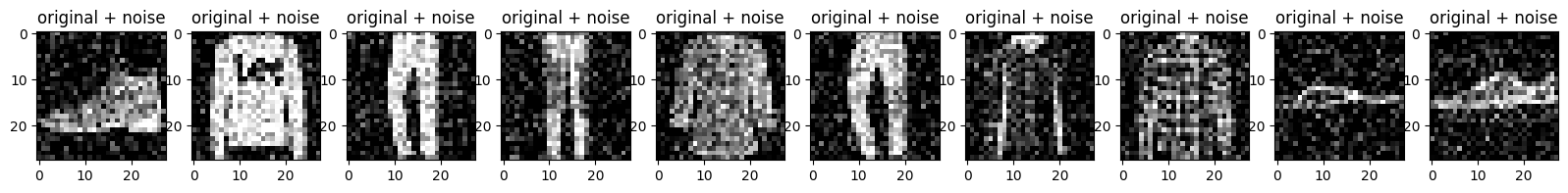
Plot the noisy images.
n = 10
plt.figure(figsize=(20, 2))
for i in range(n):
ax = plt.subplot(1, n, i + 1)
plt.title("original + noise")
plt.imshow(tf.squeeze(x_test_noisy[i]))
plt.gray()
plt.show()
Define a convolutional autoencoder
In this example, you will train a convolutional autoencoder using Conv2D layers in the encoder, and Conv2DTranspose layers in the decoder.
class Denoise(Model):
def __init__(self):
super(Denoise, self).__init__()
self.encoder = tf.keras.Sequential([
layers.Input(shape=(28, 28, 1)),
layers.Conv2D(16, (3, 3), activation='relu', padding='same', strides=2),
layers.Conv2D(8, (3, 3), activation='relu', padding='same', strides=2)])
self.decoder = tf.keras.Sequential([
layers.Conv2DTranspose(8, kernel_size=3, strides=2, activation='relu', padding='same'),
layers.Conv2DTranspose(16, kernel_size=3, strides=2, activation='relu', padding='same'),
layers.Conv2D(1, kernel_size=(3, 3), activation='sigmoid', padding='same')])
def call(self, x):
encoded = self.encoder(x)
decoded = self.decoder(encoded)
return decoded
autoencoder = Denoise()
autoencoder.compile(optimizer='adam', loss=losses.MeanSquaredError())
autoencoder.fit(x_train_noisy, x_train,
epochs=10,
shuffle=True,
validation_data=(x_test_noisy, x_test))
Epoch 1/10 1875/1875 ━━━━━━━━━━━━━━━━━━━━ 7s 2ms/step - loss: 0.0332 - val_loss: 0.0094 Epoch 2/10 1875/1875 ━━━━━━━━━━━━━━━━━━━━ 3s 2ms/step - loss: 0.0089 - val_loss: 0.0081 Epoch 3/10 1875/1875 ━━━━━━━━━━━━━━━━━━━━ 4s 2ms/step - loss: 0.0079 - val_loss: 0.0076 Epoch 4/10 1875/1875 ━━━━━━━━━━━━━━━━━━━━ 4s 2ms/step - loss: 0.0074 - val_loss: 0.0073 Epoch 5/10 1875/1875 ━━━━━━━━━━━━━━━━━━━━ 4s 2ms/step - loss: 0.0072 - val_loss: 0.0071 Epoch 6/10 1875/1875 ━━━━━━━━━━━━━━━━━━━━ 4s 2ms/step - loss: 0.0070 - val_loss: 0.0070 Epoch 7/10 1875/1875 ━━━━━━━━━━━━━━━━━━━━ 4s 2ms/step - loss: 0.0069 - val_loss: 0.0069 Epoch 8/10 1875/1875 ━━━━━━━━━━━━━━━━━━━━ 4s 2ms/step - loss: 0.0068 - val_loss: 0.0068 Epoch 9/10 1875/1875 ━━━━━━━━━━━━━━━━━━━━ 4s 2ms/step - loss: 0.0068 - val_loss: 0.0067 Epoch 10/10 1875/1875 ━━━━━━━━━━━━━━━━━━━━ 3s 2ms/step - loss: 0.0067 - val_loss: 0.0067 <keras.src.callbacks.history.History at 0x7fe6883e5a30>
Let's take a look at a summary of the encoder. Notice how the images are downsampled from 28x28 to 7x7.
autoencoder.encoder.summary()
The decoder upsamples the images back from 7x7 to 28x28.
autoencoder.decoder.summary()
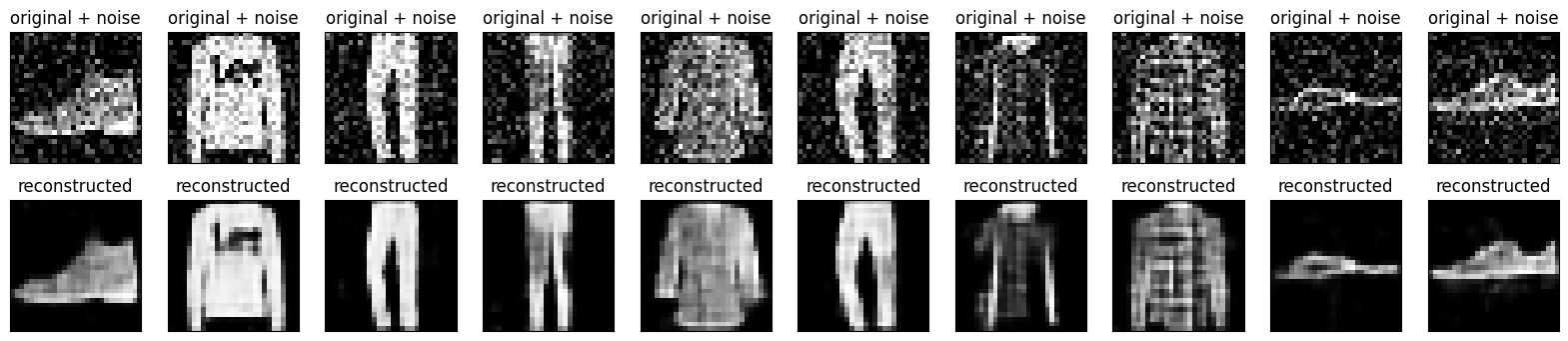
Plotting both the noisy images and the denoised images produced by the autoencoder.
encoded_imgs = autoencoder.encoder(x_test_noisy).numpy()
decoded_imgs = autoencoder.decoder(encoded_imgs).numpy()
n = 10
plt.figure(figsize=(20, 4))
for i in range(n):
# display original + noise
ax = plt.subplot(2, n, i + 1)
plt.title("original + noise")
plt.imshow(tf.squeeze(x_test_noisy[i]))
plt.gray()
ax.get_xaxis().set_visible(False)
ax.get_yaxis().set_visible(False)
# display reconstruction
bx = plt.subplot(2, n, i + n + 1)
plt.title("reconstructed")
plt.imshow(tf.squeeze(decoded_imgs[i]))
plt.gray()
bx.get_xaxis().set_visible(False)
bx.get_yaxis().set_visible(False)
plt.show()

Third example: Anomaly detection
Overview
In this example, you will train an autoencoder to detect anomalies on the ECG5000 dataset. This dataset contains 5,000 Electrocardiograms, each with 140 data points. You will use a simplified version of the dataset, where each example has been labeled either 0 (corresponding to an abnormal rhythm), or 1 (corresponding to a normal rhythm). You are interested in identifying the abnormal rhythms.
How will you detect anomalies using an autoencoder? Recall that an autoencoder is trained to minimize reconstruction error. You will train an autoencoder on the normal rhythms only, then use it to reconstruct all the data. Our hypothesis is that the abnormal rhythms will have higher reconstruction error. You will then classify a rhythm as an anomaly if the reconstruction error surpasses a fixed threshold.
Load ECG data
The dataset you will use is based on one from timeseriesclassification.com.
# Download the dataset
dataframe = pd.read_csv('http://storage.googleapis.com/download.tensorflow.org/data/ecg.csv', header=None)
raw_data = dataframe.values
dataframe.head()
# The last element contains the labels
labels = raw_data[:, -1]
# The other data points are the electrocadriogram data
data = raw_data[:, 0:-1]
train_data, test_data, train_labels, test_labels = train_test_split(
data, labels, test_size=0.2, random_state=21
)
Normalize the data to [0,1].
min_val = tf.reduce_min(train_data)
max_val = tf.reduce_max(train_data)
train_data = (train_data - min_val) / (max_val - min_val)
test_data = (test_data - min_val) / (max_val - min_val)
train_data = tf.cast(train_data, tf.float32)
test_data = tf.cast(test_data, tf.float32)
You will train the autoencoder using only the normal rhythms, which are labeled in this dataset as 1. Separate the normal rhythms from the abnormal rhythms.
train_labels = train_labels.astype(bool)
test_labels = test_labels.astype(bool)
normal_train_data = train_data[train_labels]
normal_test_data = test_data[test_labels]
anomalous_train_data = train_data[~train_labels]
anomalous_test_data = test_data[~test_labels]
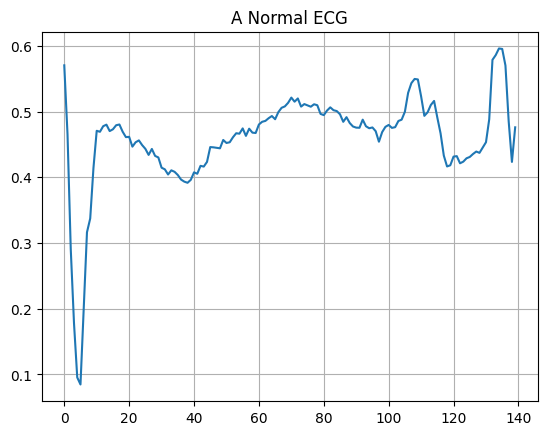
Plot a normal ECG.
plt.grid()
plt.plot(np.arange(140), normal_train_data[0])
plt.title("A Normal ECG")
plt.show()
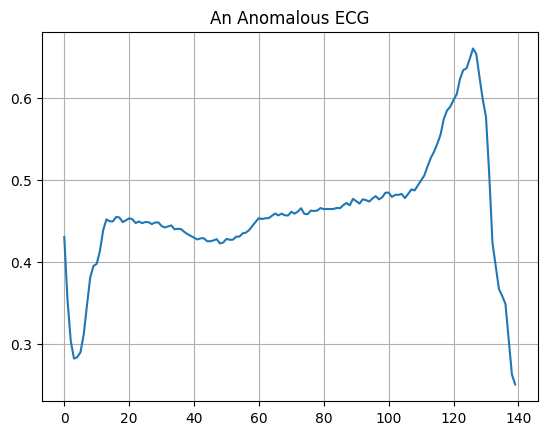
Plot an anomalous ECG.
plt.grid()
plt.plot(np.arange(140), anomalous_train_data[0])
plt.title("An Anomalous ECG")
plt.show()
Build the model
class AnomalyDetector(Model):
def __init__(self):
super(AnomalyDetector, self).__init__()
self.encoder = tf.keras.Sequential([
layers.Dense(32, activation="relu"),
layers.Dense(16, activation="relu"),
layers.Dense(8, activation="relu")])
self.decoder = tf.keras.Sequential([
layers.Dense(16, activation="relu"),
layers.Dense(32, activation="relu"),
layers.Dense(140, activation="sigmoid")])
def call(self, x):
encoded = self.encoder(x)
decoded = self.decoder(encoded)
return decoded
autoencoder = AnomalyDetector()
autoencoder.compile(optimizer='adam', loss='mae')
Notice that the autoencoder is trained using only the normal ECGs, but is evaluated using the full test set.
history = autoencoder.fit(normal_train_data, normal_train_data,
epochs=20,
batch_size=512,
validation_data=(test_data, test_data),
shuffle=True)
Epoch 1/20 5/5 ━━━━━━━━━━━━━━━━━━━━ 5s 494ms/step - loss: 0.0575 - val_loss: 0.0526 Epoch 2/20 5/5 ━━━━━━━━━━━━━━━━━━━━ 0s 14ms/step - loss: 0.0549 - val_loss: 0.0506 Epoch 3/20 5/5 ━━━━━━━━━━━━━━━━━━━━ 0s 13ms/step - loss: 0.0514 - val_loss: 0.0488 Epoch 4/20 5/5 ━━━━━━━━━━━━━━━━━━━━ 0s 13ms/step - loss: 0.0475 - val_loss: 0.0466 Epoch 5/20 5/5 ━━━━━━━━━━━━━━━━━━━━ 0s 14ms/step - loss: 0.0432 - val_loss: 0.0439 Epoch 6/20 5/5 ━━━━━━━━━━━━━━━━━━━━ 0s 13ms/step - loss: 0.0389 - val_loss: 0.0421 Epoch 7/20 5/5 ━━━━━━━━━━━━━━━━━━━━ 0s 13ms/step - loss: 0.0353 - val_loss: 0.0404 Epoch 8/20 5/5 ━━━━━━━━━━━━━━━━━━━━ 0s 13ms/step - loss: 0.0319 - val_loss: 0.0388 Epoch 9/20 5/5 ━━━━━━━━━━━━━━━━━━━━ 0s 14ms/step - loss: 0.0290 - val_loss: 0.0374 Epoch 10/20 5/5 ━━━━━━━━━━━━━━━━━━━━ 0s 13ms/step - loss: 0.0276 - val_loss: 0.0368 Epoch 11/20 5/5 ━━━━━━━━━━━━━━━━━━━━ 0s 14ms/step - loss: 0.0264 - val_loss: 0.0363 Epoch 12/20 5/5 ━━━━━━━━━━━━━━━━━━━━ 0s 14ms/step - loss: 0.0254 - val_loss: 0.0356 Epoch 13/20 5/5 ━━━━━━━━━━━━━━━━━━━━ 0s 13ms/step - loss: 0.0242 - val_loss: 0.0352 Epoch 14/20 5/5 ━━━━━━━━━━━━━━━━━━━━ 0s 14ms/step - loss: 0.0234 - val_loss: 0.0350 Epoch 15/20 5/5 ━━━━━━━━━━━━━━━━━━━━ 0s 14ms/step - loss: 0.0229 - val_loss: 0.0345 Epoch 16/20 5/5 ━━━━━━━━━━━━━━━━━━━━ 0s 14ms/step - loss: 0.0222 - val_loss: 0.0343 Epoch 17/20 5/5 ━━━━━━━━━━━━━━━━━━━━ 0s 13ms/step - loss: 0.0216 - val_loss: 0.0341 Epoch 18/20 5/5 ━━━━━━━━━━━━━━━━━━━━ 0s 14ms/step - loss: 0.0212 - val_loss: 0.0338 Epoch 19/20 5/5 ━━━━━━━━━━━━━━━━━━━━ 0s 14ms/step - loss: 0.0210 - val_loss: 0.0338 Epoch 20/20 5/5 ━━━━━━━━━━━━━━━━━━━━ 0s 14ms/step - loss: 0.0208 - val_loss: 0.0339
plt.plot(history.history["loss"], label="Training Loss")
plt.plot(history.history["val_loss"], label="Validation Loss")
plt.legend()
<matplotlib.legend.Legend at 0x7fe6704c3520>
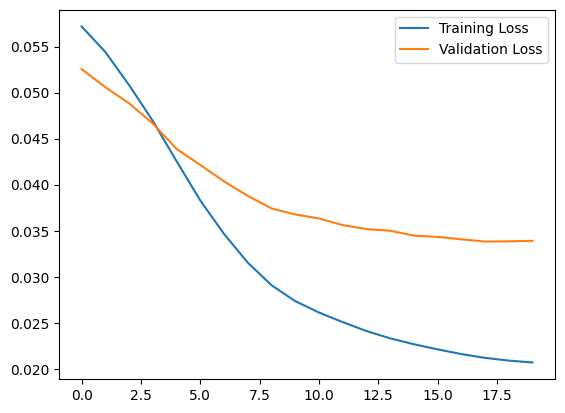
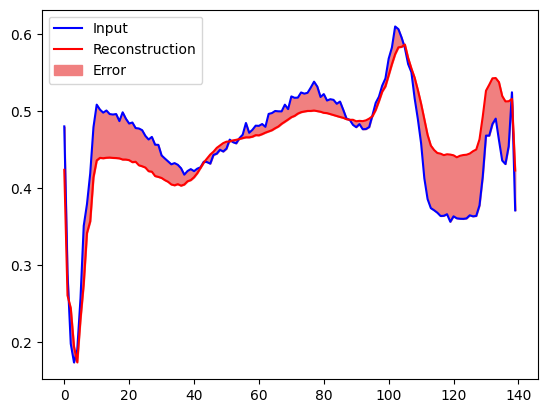
You will soon classify an ECG as anomalous if the reconstruction error is greater than one standard deviation from the normal training examples. First, let's plot a normal ECG from the training set, the reconstruction after it's encoded and decoded by the autoencoder, and the reconstruction error.
encoded_data = autoencoder.encoder(normal_test_data).numpy()
decoded_data = autoencoder.decoder(encoded_data).numpy()
plt.plot(normal_test_data[0], 'b')
plt.plot(decoded_data[0], 'r')
plt.fill_between(np.arange(140), decoded_data[0], normal_test_data[0], color='lightcoral')
plt.legend(labels=["Input", "Reconstruction", "Error"])
plt.show()
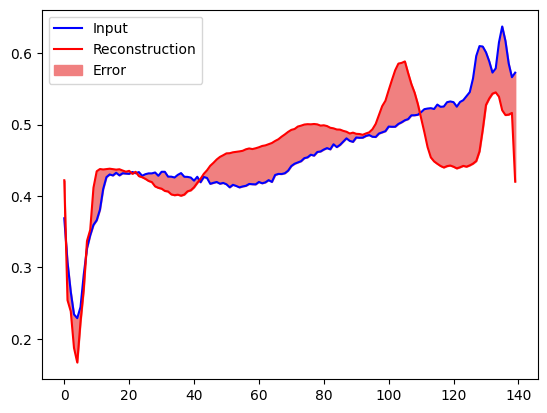
Create a similar plot, this time for an anomalous test example.
encoded_data = autoencoder.encoder(anomalous_test_data).numpy()
decoded_data = autoencoder.decoder(encoded_data).numpy()
plt.plot(anomalous_test_data[0], 'b')
plt.plot(decoded_data[0], 'r')
plt.fill_between(np.arange(140), decoded_data[0], anomalous_test_data[0], color='lightcoral')
plt.legend(labels=["Input", "Reconstruction", "Error"])
plt.show()
Detect anomalies
Detect anomalies by calculating whether the reconstruction loss is greater than a fixed threshold. In this tutorial, you will calculate the mean average error for normal examples from the training set, then classify future examples as anomalous if the reconstruction error is higher than one standard deviation from the training set.
Plot the reconstruction error on normal ECGs from the training set
reconstructions = autoencoder.predict(normal_train_data)
train_loss = tf.keras.losses.mae(reconstructions, normal_train_data)
plt.hist(train_loss[None,:], bins=50)
plt.xlabel("Train loss")
plt.ylabel("No of examples")
plt.show()
74/74 ━━━━━━━━━━━━━━━━━━━━ 1s 6ms/step
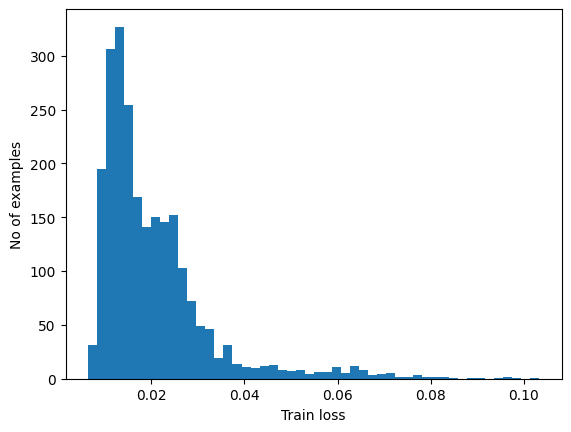
Choose a threshold value that is one standard deviations above the mean.
threshold = np.mean(train_loss) + np.std(train_loss)
print("Threshold: ", threshold)
Threshold: 0.033047937
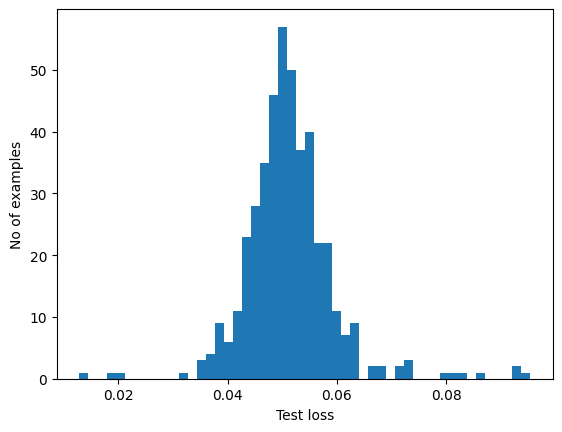
If you examine the reconstruction error for the anomalous examples in the test set, you'll notice most have greater reconstruction error than the threshold. By varing the threshold, you can adjust the precision and recall of your classifier.
reconstructions = autoencoder.predict(anomalous_test_data)
test_loss = tf.keras.losses.mae(reconstructions, anomalous_test_data)
plt.hist(test_loss[None, :], bins=50)
plt.xlabel("Test loss")
plt.ylabel("No of examples")
plt.show()
14/14 ━━━━━━━━━━━━━━━━━━━━ 0s 22ms/step
Classify an ECG as an anomaly if the reconstruction error is greater than the threshold.
def predict(model, data, threshold):
reconstructions = model(data)
loss = tf.keras.losses.mae(reconstructions, data)
return tf.math.less(loss, threshold)
def print_stats(predictions, labels):
print("Accuracy = {}".format(accuracy_score(labels, predictions)))
print("Precision = {}".format(precision_score(labels, predictions)))
print("Recall = {}".format(recall_score(labels, predictions)))
preds = predict(autoencoder, test_data, threshold)
print_stats(preds, test_labels)
Accuracy = 0.944 Precision = 0.9921875 Recall = 0.9071428571428571
Next steps
To learn more about anomaly detection with autoencoders, check out this excellent interactive example built with TensorFlow.js by Victor Dibia. For a real-world use case, you can learn how Airbus Detects Anomalies in ISS Telemetry Data using TensorFlow. To learn more about the basics, consider reading this blog post by François Chollet. For more details, check out chapter 14 from Deep Learning by Ian Goodfellow, Yoshua Bengio, and Aaron Courville.
